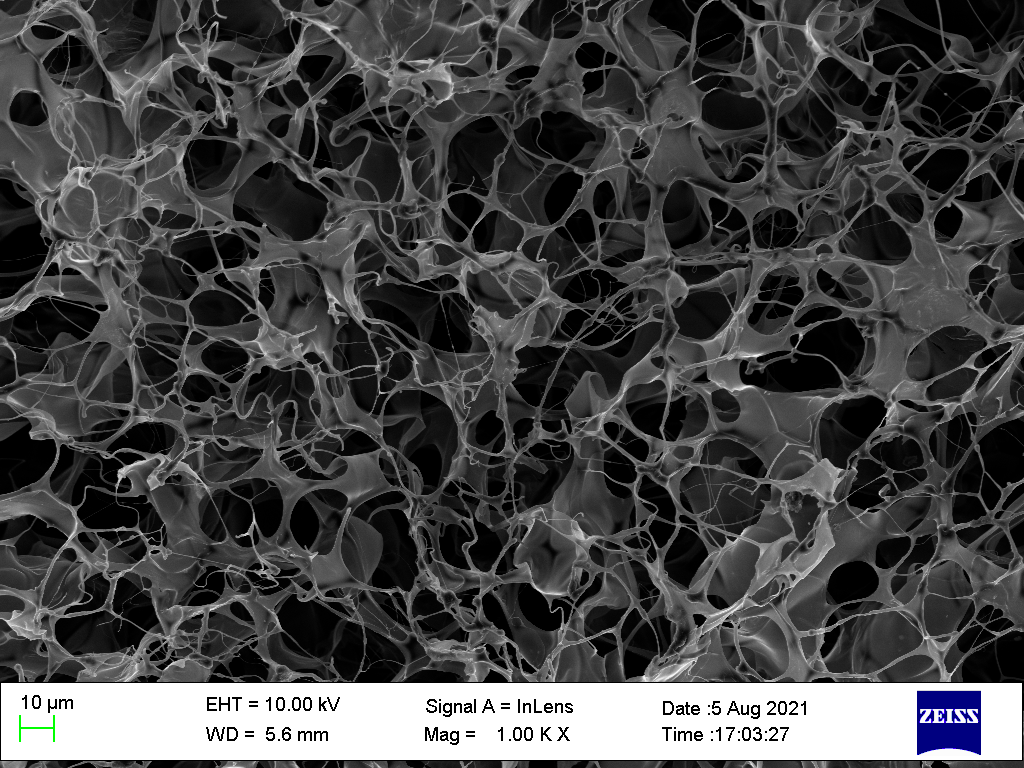
porous cryogel
This was a biomaterials project I reproduced during my undergraduate
studies, based on the injectable shape-memory scaffold reported by
Bencherif and colleagues. The concept was simple but elegant: to create
a porous hydrogel that could be compressed, injected through a syringe,
and then recover its original structure after injection.
In their work, the porous architecture was generated through slow
covalent crosslinking at low temperature. Ice crystals formed during the
crosslinking process occupied space within the material and later served
as pore templates.
Later, my colleagues and I developed a related approach for producing an
ionically crosslinked porous alginate hydrogel. We first froze and
lyophilised the alginate precursor, then immersed it in a calcium-ion
solution to induce crosslinking. This produced a highly similar porous
structure, while also preserving the material’s injectable behaviour.
Our original aim was to load NK cells into the hydrogel and use it as a
delivery platform for solid tumours. Because of COVID and a few other
unavoidable disruptions, I never managed to complete the bioengineering
part of the project. It was a slightly frustrating ending.
Reference: Sidi A. Bencherif, R. Warren Sands, Deen
Bhatta, et al. “Injectable preformed scaffolds with shape-memory
properties.” Proceedings of the National Academy of Sciences
109(48): 19590-19595 (2012).
doi.org/10.1073/pnas.1211516109